Documentation#

GAUCHE is a collaborative, open-source software library that aims to make state-of-the-art probabilistic modelling and black-box optimisation techniques more easily accessible to scientific experts in chemistry, materials science and beyond. We provide 30+ bespoke kernels for molecules, chemical reactions and proteins and illustrate how they can be used for Gaussian processes and Bayesian optimisation in 10+ easy-to-adapt tutorial notebooks.
Overview#
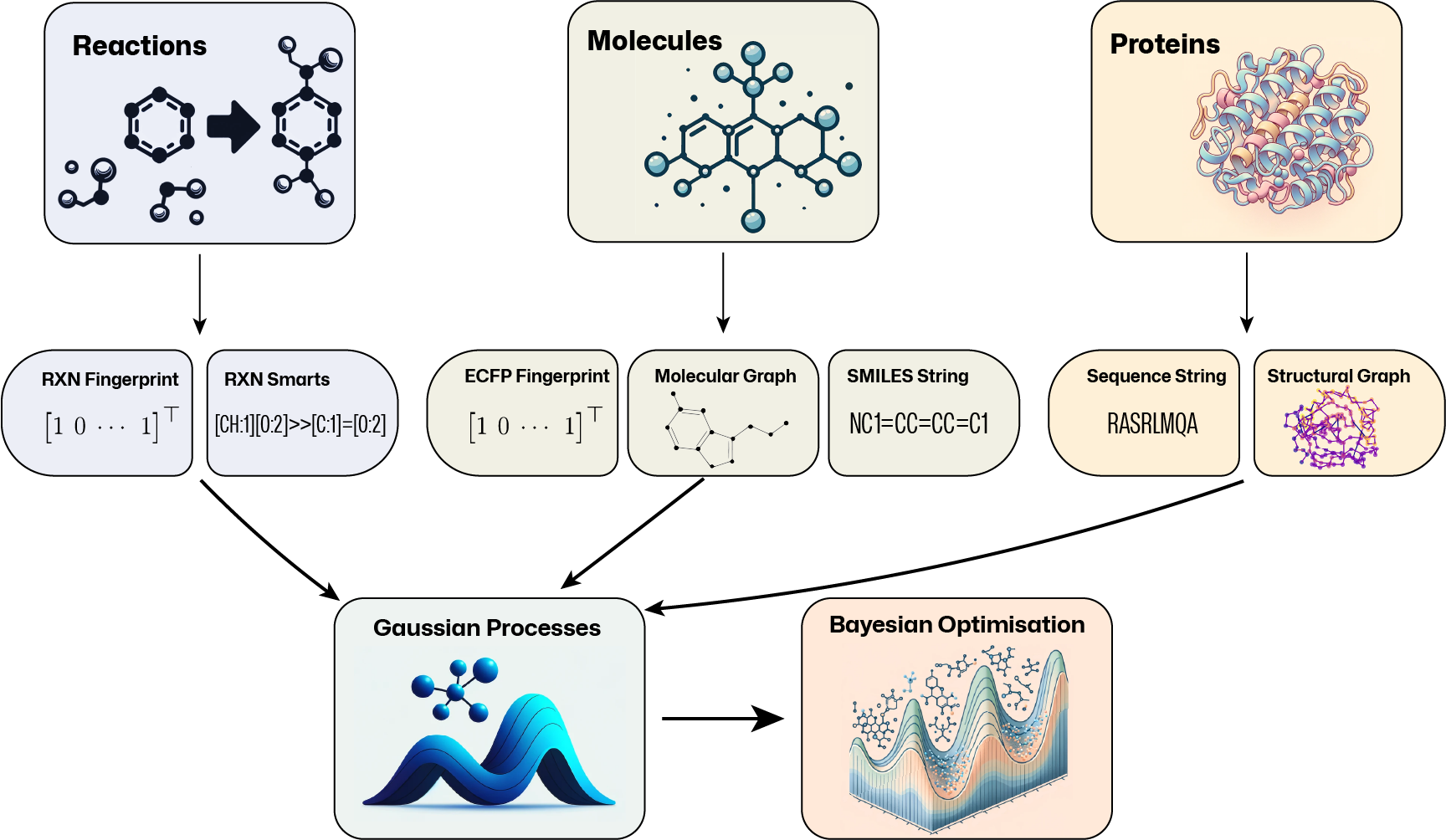
General-purpose Gaussian process (GP) and Bayesian optimisation (BO) libraries do not cater for molecular representations. Likewise, general-purpose molecular machine learning libraries do not consider GPs and BO. To bridge this gap, GAUCHE provides a modular, robust and easy-to-use framework of 30+ parallelisable and batch-GP-compatible implementations of string, fingerprint and graph kernels that operate on a range of widely-used molecular representations.
Kernels#
Standard GP packages typically assume continuous input spaces of low and fixed dimensionality. This makes it difficult to apply them to common molecular representations: molecular graphs are discrete objects, SMILES strings vary in length and topological fingerprints tend to be high-dimensional and sparse. To bridge this gap, GAUCHE provides:
Fingerprint Kernels that measure the similarity between bit/count vectors of descriptor by examining the degree to which their elements overlap.
String Kernels that measure the similarity between strings by examining the degree to which their sub-strings overlap.
Graph Kernels that measure between graphs by examining the degree to which certain substructural motifs overlap.
Representations#
GAUCHE supports any representation that is based on bit/count vectors, strings or graphs. For rapid prototyping and benchmarking, we also provide a range of standard featurisation techniques for molecules, chemical reactions and proteins:
Domain |
Representation |
|---|---|
Molecules |
ECFP Fingerprints [1], rdkit Fragments, Fragprints, Graphs [2], SMILES [3], SELFIES [4] |
Chemical Reactions |
One-Hot Encoding, Data-Driven Reaction Fingerprints [5], Differential Reaction Fingerprints [6], Reaction SMARTS |
Proteins |
Sequences, Graphs [2] |
Getting Started#
The easiest way to install Gauche is via pip.
pip install gauche
As not all users will need the full functionality of the package, we provide a range of installation options:
pip install gauche - installs the core functionality of GAUCHE (kernels, representations, data loaders, etc.) and should cover a wide range of use cases.
pip install gauche[rxn] - additionally installs the rxnfp and drfp fingerprints that can be used to represent chemical reactions.
pip install gauche[graphs] - installs all dependencies for graph kernels and representations.
If you aren’t sure which installation option is right for you, you can simply install all of them with pip install gauche[all].
The best way to get started with GAUCHE is to check out our tutorial notebooks. These notebooks provide a step-by-step introduction to the core functionality of GAUCHE and illustrate how it can be used to solve a range of common problems in molecular property prediction and optimisation.
- Loading and Featurising Molecular Data
- GP Regression on Molecules
- Bayesian Optimisation Over Molecules
- Sparse GP Regression on Molecules
- Multitask GP Regression on Molecules
- Learning an Objective Function through Interaction with a Human Chemist
- Preferential Bayesian Optimisation
- GP Regression on Protein Sequences: Bag of Amino Acids
- GP Regression on Protein Sequences: Subsequence String Kernel
- Bayesian GNNs for Molecular Property Prediction
Extensions#
If there are any specific kernels or representations that you would like to see included in GAUCHE, please reach out or submit an issue/pull request.
Gauche’s API#
API Reference
- gauche.kernels
- Fingerprint Kernels
TanimotoKernelbatch_tanimoto_sim()BraunBlanquetKernelbatch_braun_blanquet_sim()DiceKernelbatch_dice_sim()FaithKernelbatch_faith_sim()ForbesKernelbatch_forbes_sim()InnerProductKernelbatch_inner_product_sim()IntersectionKernelbatch_intersection_sim()MinMaxKernelbatch_minmax_sim()OtsukaKernelbatch_otsuka_sim()RandKernelbatch_rand_sim()RogersTanimotoKernelbatch_rogers_tanimoto_sim()RussellRaoKernelbatch_russell_rao_sim()SogenfreiKernelbatch_sogenfrei_sim()SokalSneathKernelbatch_sokal_sneath_sim()
- Graph Kernels
- String Kernels
- Fingerprint Kernels
- gauche.representations
- gauche.dataloader
Indices and tables#
References#
[1] Rogers, D. and Hahn, M., 2010. Extended-connectivity fingerprints. Journal of Chemical Information and Modeling, 50(5), pp.742-754.
[2] Jamasb, A., Viñas Torné, R., Ma, E., Du, Y., Harris, C., Huang, K., Hall, D., Lió, P. and Blundell, T., 2022. Graphein-a Python library for geometric deep learning and network analysis on biomolecular structures and interaction networks. Advances in Neural Information Processing Systems, 35, pp.27153-27167.
[3] Weininger, D., 1988. SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. Journal of Chemical Information and Computer Sciences, 28(1), pp.31-36.
[4] Krenn, M., Häse, F., Nigam, A., Friederich, P. and Aspuru-Guzik, A., 2020. Self-referencing embedded strings (SELFIES): A 100% robust molecular string representation. Machine Learning: Science and Technology, 1(4), p.045024.
[5] Probst, D., Schwaller, P. and Reymond, J.L., 2022. Reaction classification and yield prediction using the differential reaction fingerprint DRFP. Digital Discovery, 1(2), pp.91-97.
[6] Schwaller, P., Probst, D., Vaucher, A.C., Nair, V.H., Kreutter, D., Laino, T. and Reymond, J.L., 2021. Mapping the space of chemical reactions using attention-based neural networks. Nature Machine Intelligence, 3(2), pp.144-152.